#Researchers develop new cytosine base editors with high specificity and precision
“#Researchers develop new cytosine base editors with high specificity and precision”
Base editors, which enable production of highly efficient targeted point mutations in genomic DNA without causing double-stranded DNA breaks, hold great promise for gene therapy in human disease and trait improvement in crop plants.
Gao Caixia’s group from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences has been developing plant base editing technologies. The researchers showed that cytosine base editors (CBEs) induce unexpected genome-wide off-target mutations in rice in order to stimulate the worldwide genome editing community to correct them.
Recently, GAO’s team created two new CBEs based on a truncated human APOBEC3 cytidine deaminase (A3Bctd) and developed a high-throughput assay for assessing sgRNA-independent deamination changes in plant CBEs.
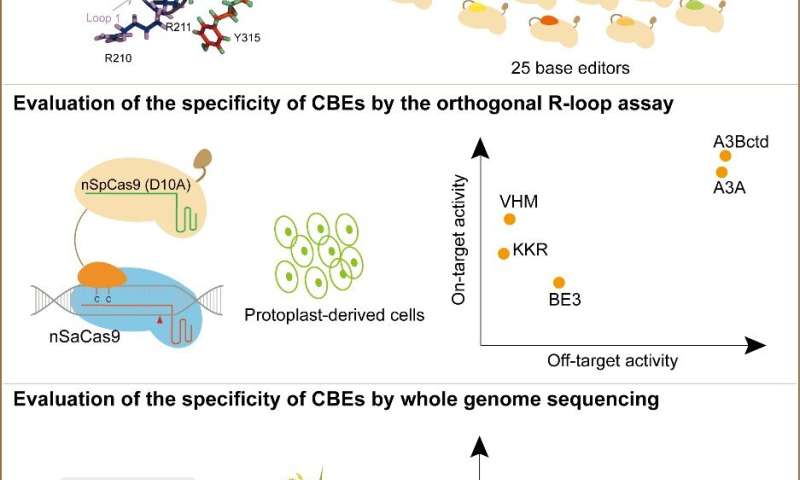
They first developed a rapid, high-throughput and inexpensive method for assessing CBEs in plants (nSaCas9-mediated orthogonal R-loop assay). In this assay, the orthogonal CRISPR system, nSaCas9, was used to create ssDNA regions in plant cells that acted as targets for sgRNA-independent deamination changes. To assess the nSaCas9-mediated orthogonal R-loop assay, they compared it with the whole-genome sequencing (WGS) assay.
The consistent results indicate that the nSaCas9-mediated orthogonal R-loop assay provides a logical, rapid and high-throughput method for assessing the sgRNA-independent off-target activities of CBEs.
Then they created 16 A3Bctd deaminase variants by rational design and evaluated their on-target efficiency and the sgRNA-independent off-target activities. They tested these A3Bctd-BE3 variants using the R-loop assay and selected seven mutations associated with efficient on-target editing activity and reduced off-target activity. After that, they combined these mutations and produced nine new A3Bctd-BE3 variants with double or triple amino acid substitutions.
In this way, they obtained two new CBE variants, A3Bctd-VHM-BE3 and A3Bctd-KKR-BE3, which exhibited efficient on-target activity and markedly reduced sgRNA-independent off-target activity.
In addition, these two new CBEs behaved more precisely at their target sites, and mainly produced single and double C edits. Also, they validated the high specificity of A3Bctd-VHM-BE3 and A3Bctd-KKR-BE3 by WGS assay.
The scientific paper, entitled “Rationally designed APOBEC3B cytosine base editors with improved specificity,” was online published in Molecular Cell on July 27.
The research was mainly supported by the National Transgenic Science and Technology Program, the Strategic Priority Research Program of the Chinese Academy of Sciences, and the National Natural Science Foundation of China.
More information:
Shuai Jin et al. Rationally Designed APOBEC3B Cytosine Base Editors with Improved Specificity, Molecular Cell (2020). DOI: 10.1016/j.molcel.2020.07.005
Researchers develop new cytosine base editors with high specificity and precision (2020, July 27)
retrieved 27 July 2020
from https://phys.org/news/2020-07-cytosine-base-editors-high-specificity.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.
If you want to read more Like this articles, you can visit our Science category.
if you want to watch Movies or Tv Shows go to Dizi.BuradaBiliyorum.Com for forums sites go to Forum.BuradaBiliyorum.Com



